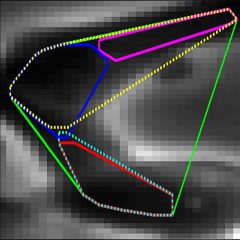
LEVER (Lineage Editing and Validation) is a collection of software tools for analyzing the development of proliferating (dividing) cells in time-lapse microscopy image sequences. LEVER includes algorithms for segmentation, tracking and lineaging. LEVER also includes a user interface that allows the segmentation, tracking and lineaging results to be validated, with any errors in the automated processing easily identified and quickly corrected
The key to analyzing time-lapse microscopy images of live proliferating cells is the segmentation algorithm. Given perfect segmentation and sufficient temporal resolution of imaging, the tracking and lineaging algorithms will never make a mistake. Sadly, (or happily if you like writing segmentation algorithms) perfect segmentation will never exist. The entire LEVER architecture is designed to use temporal data from the tracking and contextual information on spatio-temporal population dynamics from the lineaging to improve the performance of the segmentation algorithm.
Versions of LEVER
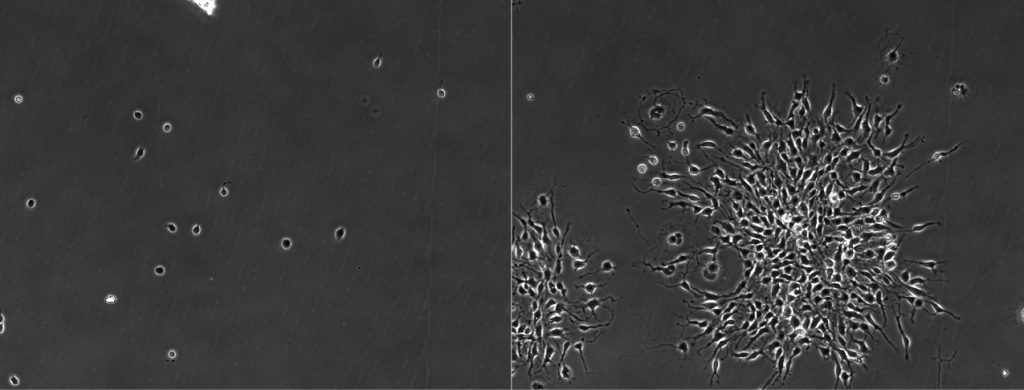
The version of LEVER available here includes segmentation algorithms for adult and embryonic mouse neural stem cells imaged with phase contrast microscopy. There are as yet un-released segmentation algorithms for phase/brightfield and FUCCI images of human cancer cells and mouse T cells. There are also versions of LEVER that include 3-D visualization and segmentation for 5-D (3-D+time+channels) fluorescence microscopy including confocal, multi-photon and lattice light sheet. These algorithms will generally be released concurrent with the publication of the related manuscript(s). We are always interested in developing new collaborations – please contact acohen ‘at’ coe.drexel.edu if you are interested in working with us to develop a LEVER segmentation for your application.
Using LEVER
Here you can read about how to:
- start LEVER
- import images and metadata into LEVER’s format
- draw a lineage tree
- resegment from a lineage tree
- lock and freeze your processed data
- write a custom segmentation script